DEanalysis_bulkRNA
Anthony Hung
2021-01-19
Last updated: 2021-02-22
Checks: 7 0
Knit directory: invitroOA_pilot_repository/
This reproducible R Markdown analysis was created with workflowr (version 1.6.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20210119) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 3500638. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: code/bulkRNA_preprocessing/.snakemake/conda-archive/
Ignored: code/bulkRNA_preprocessing/.snakemake/conda/
Ignored: code/bulkRNA_preprocessing/.snakemake/locks/
Ignored: code/bulkRNA_preprocessing/.snakemake/shadow/
Ignored: code/bulkRNA_preprocessing/.snakemake/singularity/
Ignored: code/bulkRNA_preprocessing/.snakemake/tmp.3ekfs3n5/
Ignored: code/bulkRNA_preprocessing/fastq/
Ignored: code/bulkRNA_preprocessing/out/
Ignored: code/single_cell_preprocessing/.snakemake/conda-archive/
Ignored: code/single_cell_preprocessing/.snakemake/conda/
Ignored: code/single_cell_preprocessing/.snakemake/locks/
Ignored: code/single_cell_preprocessing/.snakemake/shadow/
Ignored: code/single_cell_preprocessing/.snakemake/singularity/
Ignored: code/single_cell_preprocessing/YG-AH-2S-ANT-1_S1_L008/
Ignored: code/single_cell_preprocessing/YG-AH-2S-ANT-2_S2_L008/
Ignored: code/single_cell_preprocessing/demuxlet/.DS_Store
Ignored: code/single_cell_preprocessing/fastq/
Ignored: data/external_scRNA/Chou_et_al2020/
Ignored: data/external_scRNA/Jietal2018/
Ignored: data/external_scRNA/Wuetal2021/
Ignored: data/external_scRNA/merged_external_scRNA.rds
Ignored: data/poweranalysis/alasoo_et_al/
Ignored: output/GO_terms_enriched.csv
Ignored: output/topicModel_k=6.rds
Ignored: output/topicModel_k=7.rds
Ignored: output/topicModel_k=8.rds
Ignored: output/voom_results.rds
Unstaged changes:
Modified: .gitignore
Deleted: analysis/figure/topicModel_scRNA.Rmd/plot correlations external-1.png
Deleted: analysis/figure/topicModel_scRNA.Rmd/structure plot external-1.png
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were made to the R Markdown (analysis/DEanalysis_bulkRNA.Rmd) and HTML (docs/DEanalysis_bulkRNA.html) files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view the files as they were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | 77338dc | Anthony Hung | 2021-01-21 | knit analysis files |
| Rmd | 5026544 | Anthony Hung | 2021-01-21 | permute 10 times |
| html | 5026544 | Anthony Hung | 2021-01-21 | permute 10 times |
| Rmd | 8542407 | Anthony Hung | 2021-01-21 | knit anlaysis files |
| html | 8542407 | Anthony Hung | 2021-01-21 | knit anlaysis files |
| Rmd | 78cfbcd | Anthony Hung | 2021-01-21 | finish paring down files |
| Rmd | e3fbd0f | Anthony Hung | 2021-01-21 | add ward et al data files to data |
| Rmd | 0711682 | Anthony Hung | 2021-01-21 | start normalization file |
| Rmd | 28f57fa | Anthony Hung | 2021-01-19 | Add files for analysis |
Introduction
This code takes as input the filtered count data and two factors of unwanted variation fit by RUVs. It then uses limma and voom to perform differential expression analysis between treatment conditions using the data. After we have a set of differentially expressed genes, we look for enrichments amongst DE genes in GO pathways and previously published osteoarthritis datasets.
Load libraries and data
library("limma")
library("plyr")
library("dplyr")
library("edgeR")
library("tidyr")
library("ashr")
library("ggplot2")
library("RUVSeq")
library("topGO")
#load in filtered count data, RUVs output
filt_counts <- readRDS("data/filtered_counts.rds")
filt_counts <- filt_counts$counts
RUVsOut <- readRDS("data/RUVsOut.rds")
# load gene annotations
gene_anno <- read.delim("data/gene-annotation.txt",
sep = "\t")
# load in reordered sample information
sampleinfo <- readRDS("data/Sample.info.RNAseq.reordered.rds")Specify design matrix for Linear mixed model
anno <- pData(RUVsOut)
x <- paste0(anno$Individual, anno$treatment)
anno$LibraryPrepBatch <- factor(anno$LibraryPrepBatch, levels = c("1", "2"))
anno$Replicate <- factor(anno$Replicate, levels = c("1", "2", "3"))
design <- model.matrix(~treatment + Individual + W_1 + W_2 + RIN,
data = anno)
colnames(design) <- gsub("treatment", "", colnames(design))
colnames(design)[1] "(Intercept)" "Unstrain" "IndividualNA18856 "
[4] "IndividualNA19160 " "W_1" "W_2"
[7] "RIN" Fit the model using limma and voom
# Model replicate as a random effect
# Recommended to run both voom and duplicateCorrelation twice.
# https://support.bioconductor.org/p/59700/#67620
#TMM Normalization
y <- DGEList(filt_counts)
y <- calcNormFactors(y, method = "TMM")
#Voom for differential expression
v1 <- voom(y, design)
corfit1 <- duplicateCorrelation(v1, design, block = x)Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievablecorfit1$consensus.correlation[1] 0.3550961v2 <- voom(y, design, block = x, correlation = corfit1$consensus.correlation)
corfit2 <- duplicateCorrelation(v2, design, block = x)Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievablecorfit2$consensus.correlation[1] 0.3552816fit <- lmFit(v2, design, block = x,
correlation = corfit2$consensus.correlation)
fit <- eBayes(fit)
saveRDS(v2, "output/voom_results.rds")Assess model results from voom
#this function extracts the results from the voom output for each of the variables (each "test")
get_results <- function(x, number = nrow(x$coefficients), sort.by = "none",
...) {
# x - object MArrayLM from eBayes output
# ... - additional arguments passed to topTable
stopifnot(class(x) == "MArrayLM")
results <- topTable(x, number = number, sort.by = sort.by, ...)
return(results)
}
results_treatment <- get_results(fit, coef = "Unstrain",
number = nrow(filt_counts), sort = "none")
ma_treatment <- ggplot(data.frame(Amean = fit$Amean, logFC = fit$coef[, "Unstrain"]),
aes(x = Amean, y = logFC)) +
geom_point() +
labs(x = "Average expression level", y = "Log fold change",
title = "Treatment effect")
ma_treatment
| Version | Author | Date |
|---|---|---|
| 8542407 | Anthony Hung | 2021-01-21 |
hist_treatment <- ggplot(results_treatment, aes(x = P.Value)) +
geom_histogram(binwidth = 0.01) +
labs(x = "p-value", y = "Number of genes", title = "Treatment effect")
hist_treatment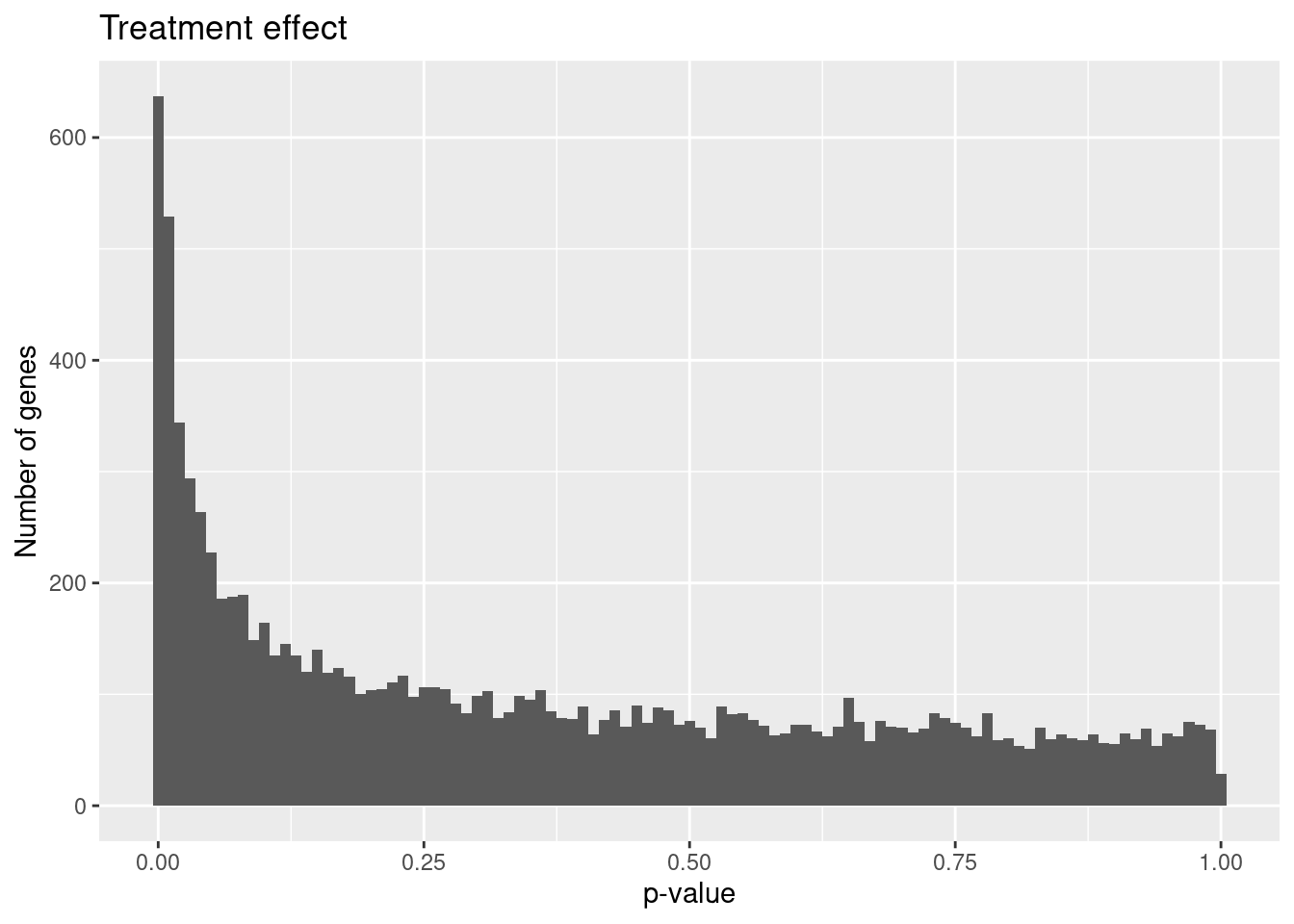
| Version | Author | Date |
|---|---|---|
| 8542407 | Anthony Hung | 2021-01-21 |
Use ash for mutliple testing correction
Run ash on the treatment results.
run_ash <- function(x, coef) {
# Perform multiple testing correction with adaptive shrinkage (ASH)
#
# x - object MArrayLM from eBayes output
# coef - coefficient tested by eBayes
stopifnot(class(x) == "MArrayLM", coef %in% colnames(x$coefficients))
result <- ash(betahat = x$coefficients[, coef],
sebetahat = x$stdev.unscaled[, coef] * sqrt(x$s2.post),
df = x$df.total[1])
return(result)
}
ash_treatment <- run_ash(fit, "Unstrain")
class(ash_treatment)[1] "ash"names(ash_treatment)[1] "fitted_g" "loglik" "logLR" "data" "result" "call" sum(ash_treatment$result$svalue < .05)[1] 773hist(ash_treatment$result$svalue)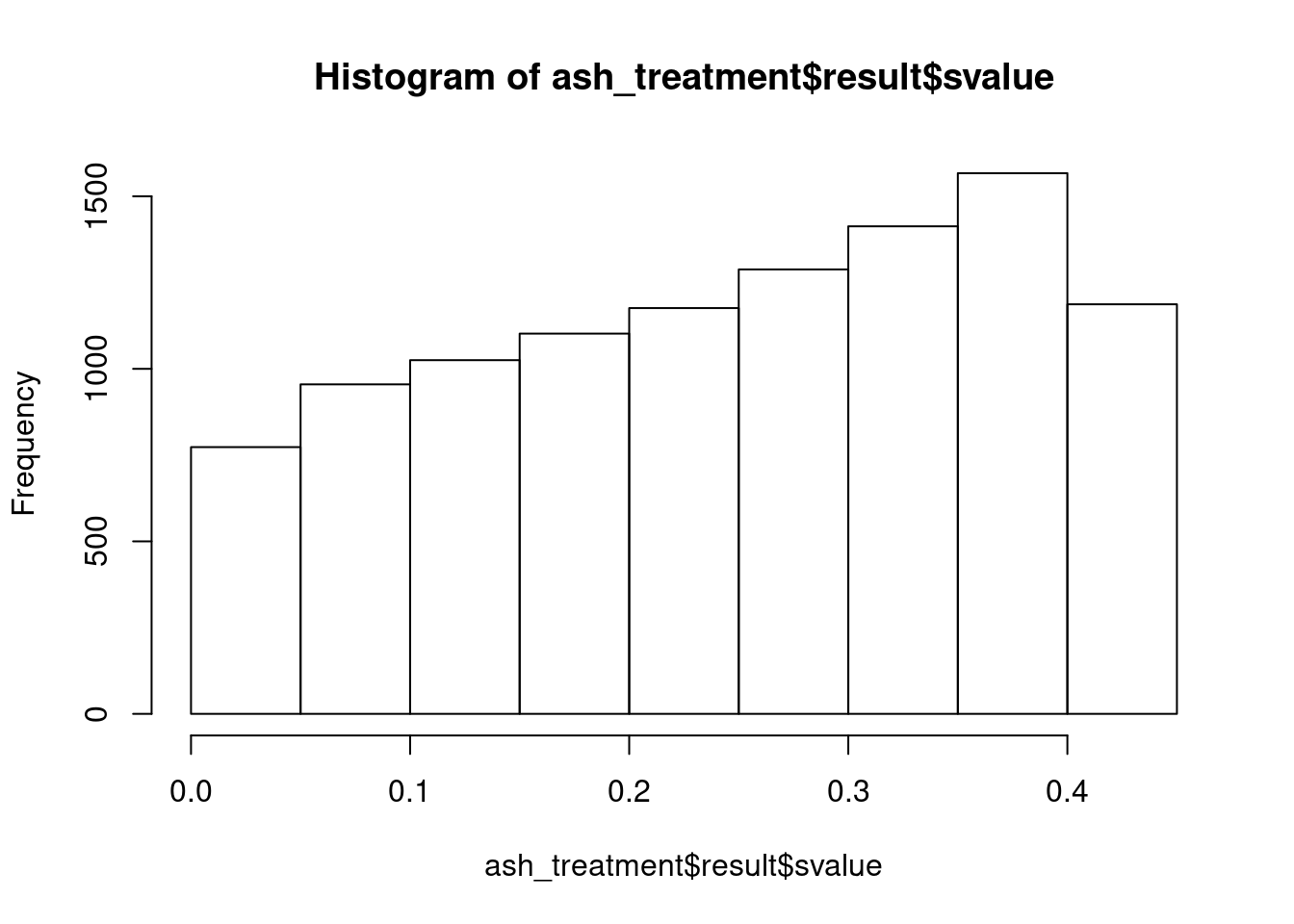
| Version | Author | Date |
|---|---|---|
| 8542407 | Anthony Hung | 2021-01-21 |
Plots
Functions to make plots from the results
plot_ma <- function(x, qval) {
# Create MA plot.
#
# x - data frame with topTable and ASH output
# (columns logFC, AveExpr, and qvalue)
# qval - qvalue cutoff for calling a gene DE
#
stopifnot(is.data.frame(x), c("logFC", "AveExpr", "qvalue") %in% colnames(x),
is.numeric(qval), qval <= 1, qval >= 0)
x$highlight <- ifelse(x$qvalue < qval, "darkred", "gray75")
x$highlight <- factor(x$highlight, levels = c("darkred", "gray75"))
ggplot(x, aes(x = AveExpr, y = logFC, color = highlight, shape = highlight)) +
geom_point() +
labs(x = "Average expression level", y = "Log fold change") +
scale_color_identity(drop = FALSE) +
scale_shape_manual(values = c(16, 1), drop = FALSE) +
theme(legend.position = "none")
# scale_color_gradient(low = "red", high = "white", limits = c(0, 0.25))
}
plot_volcano <- function(x, qval) {
# Create volcano plot.
#
# x - data frame with topTable and ASH output
# (columns logFC, P.Value, and qvalue)
# qval - qvalue cutoff for calling a gene DE
#
stopifnot(is.data.frame(x), c("logFC", "P.Value", "qvalue") %in% colnames(x),
is.numeric(qval), qval <= 1, qval >= 0)
x$highlight <- ifelse(x$qvalue < qval, "darkred", "gray75")
x$highlight <- factor(x$highlight, levels = c("darkred", "gray75"))
ggplot(x, aes(x = logFC, y = -log10(P.Value), color = highlight)) +
geom_point(shape = 1) +
labs(x = "Log fold change",
y = expression(-log[10] * " p-value")) +
scale_color_identity(drop = FALSE) +
theme(legend.position = "none")
}
plot_pval_hist <- function(x, qval) {
# Create histogram of p-values.
#
# x - data frame with topTable and ash output (columns P.Value and qvalue)
# qval - qvalue cutoff for calling a gene DE
#
stopifnot(is.data.frame(x), c("P.Value", "qvalue") %in% colnames(x))
x$highlight <- ifelse(x$qvalue < qval, "darkred", "gray75")
x$highlight <- factor(x$highlight, levels = c("darkred", "gray75"))
ggplot(x, aes(x = P.Value, fill = highlight)) +
geom_histogram(position = "stack", binwidth = 0.01) +
scale_fill_identity(drop = FALSE) +
labs(x = "p-value", y = "Number of genes")
}Apply functions to build plots for FDR 0.05
tests <- colnames(fit$coefficients)
results <- vector(length = length(tests), mode = "list")
names(results) <- tests
for (test in tests[c(1:7)]) {
# Extract limma results
print(test)
results[[test]] <- get_results(fit, coef = test)
# Add mutliple testing correction with ASH
output_ash <- run_ash(fit, coef = test)$result
results[[test]] <- cbind(results[[test]], lfsr = output_ash$lfsr,
lfdr = output_ash$lfdr, qvalue = output_ash$qvalue,
svalue = output_ash$svalue)
}[1] "(Intercept)"
[1] "Unstrain"
[1] "IndividualNA18856 "
[1] "IndividualNA19160 "
[1] "W_1"
[1] "W_2"
[1] "RIN"#FDR 0.05
plot_ma(results[["Unstrain"]], 0.05)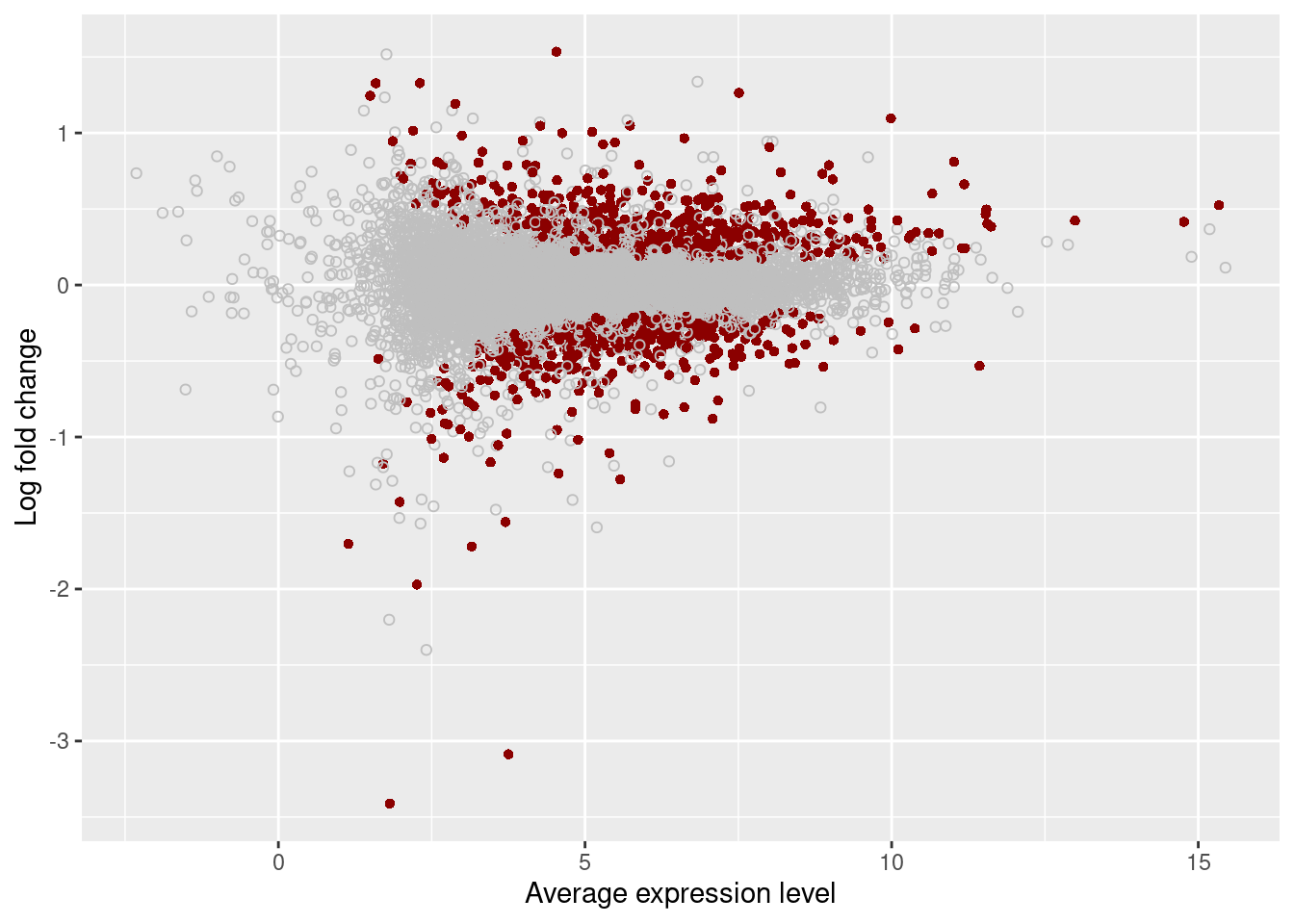
| Version | Author | Date |
|---|---|---|
| 8542407 | Anthony Hung | 2021-01-21 |
plot_volcano(results[["Unstrain"]], 0.05)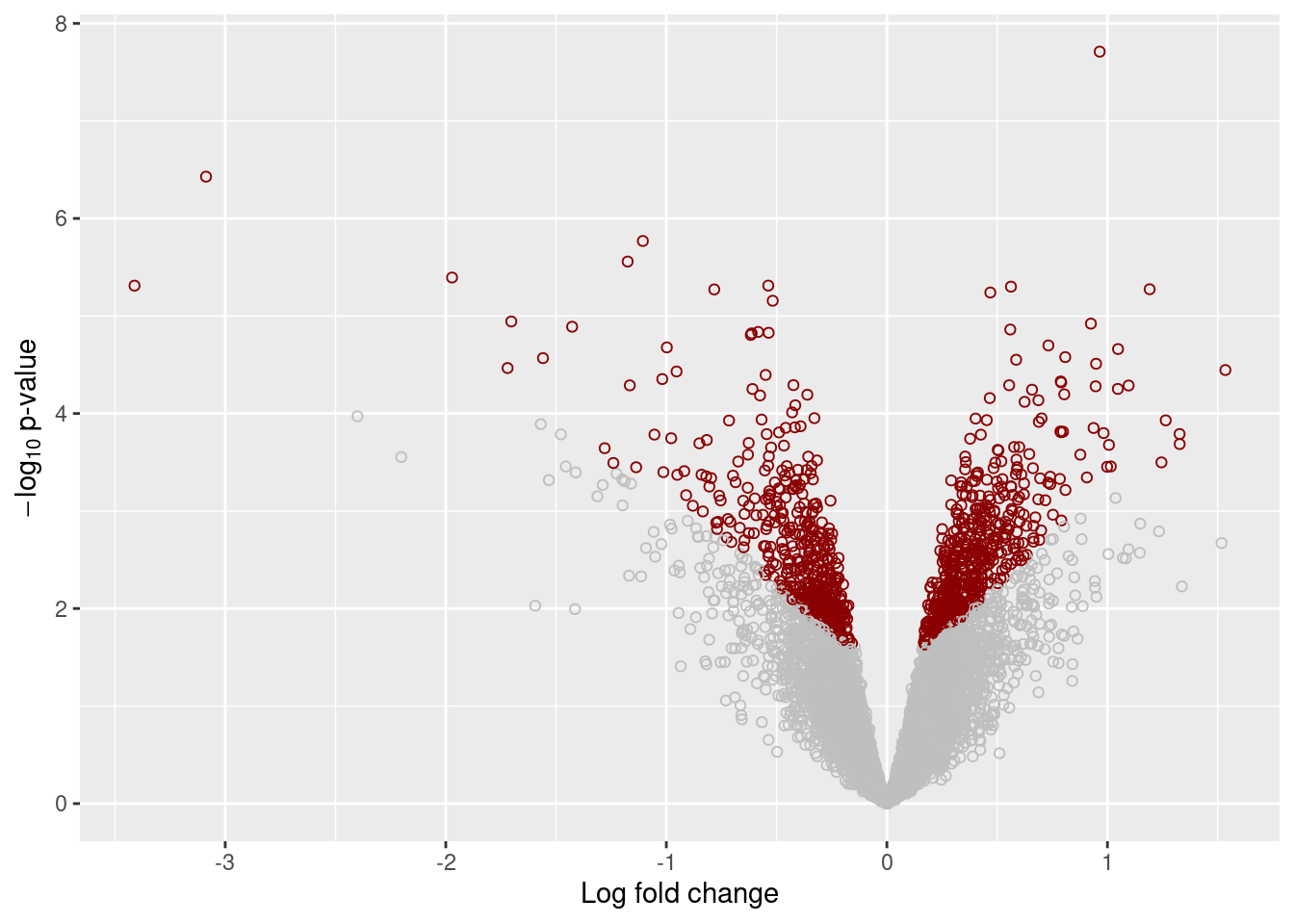
| Version | Author | Date |
|---|---|---|
| 8542407 | Anthony Hung | 2021-01-21 |
plot_pval_hist(results[["Unstrain"]], 0.05)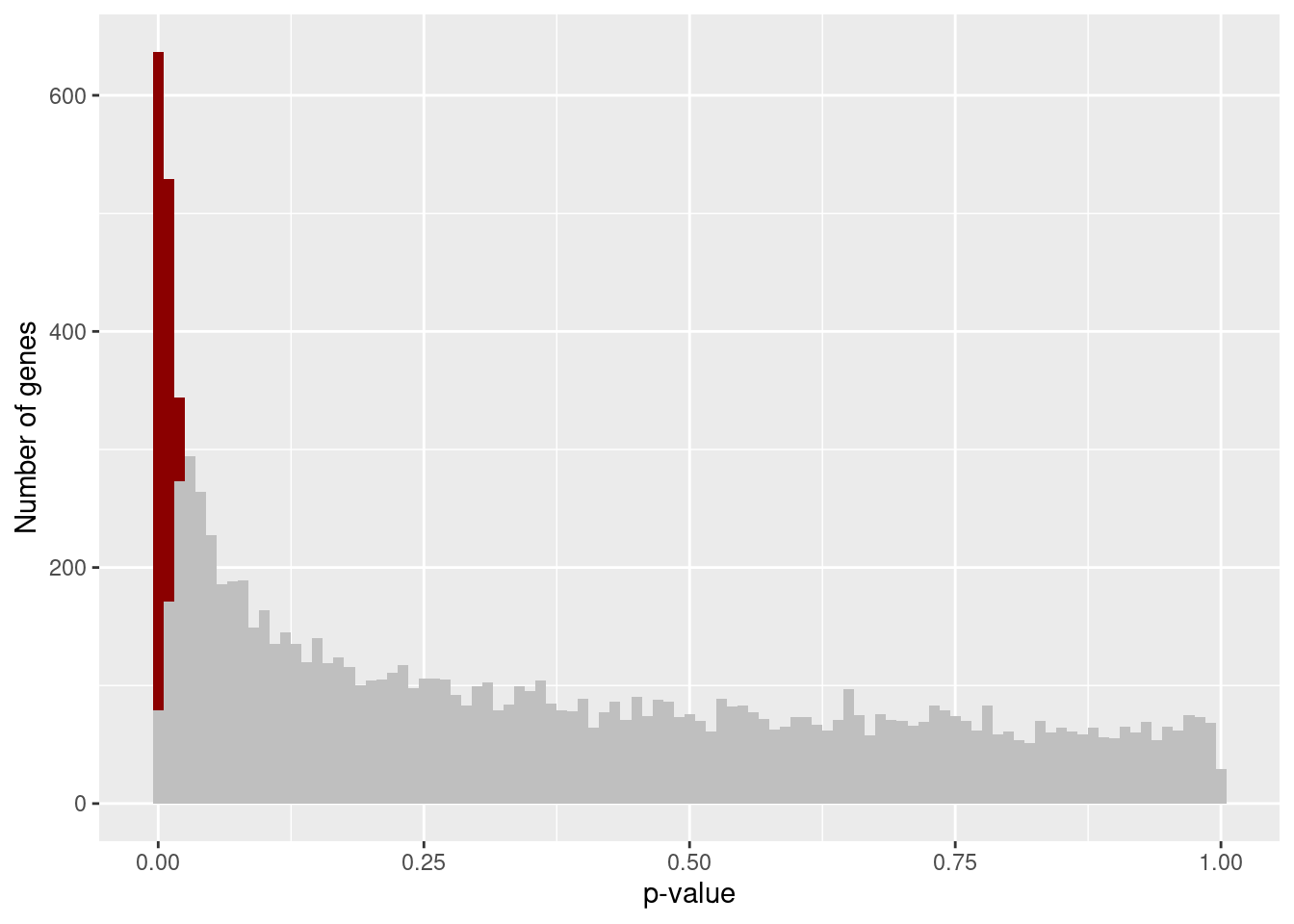
| Version | Author | Date |
|---|---|---|
| 8542407 | Anthony Hung | 2021-01-21 |
table(results[["Unstrain"]]$qvalue < 0.05)
FALSE TRUE
9499 987 significant_genes_05 <- row.names(results[["Unstrain"]])[results[["Unstrain"]]$qvalue < 0.05]
significant_symbols_05 <- gene_anno$external_gene_name[match(significant_genes_05, gene_anno$ensembl_gene_id)]
head(significant_symbols_05)[1] DVL1 MRPL20 MMP23B PEX10 CEP104 ACOT7
56858 Levels: A1BG A1BG-AS1 A1CF A2M A2M-AS1 A2ML1 A2ML1-AS1 ... ZZEF1significant_anno_05 <- gene_anno[match(significant_genes_05, gene_anno$ensembl_gene_id),]
DE_results_original <- resultsGene ontology analysis with topGO
Use topGO for GO analysis. It accounts for the nested graph structure of GO terms to prune the number of GO categories tested (Alexa et al. 2006). Essentially, it decreases the redundancy of the results.
# k-s test making use of ranked based on p-values
genes <- results[["Unstrain"]]$qvalue
names(genes) <- rownames(results[["Unstrain"]])
selection <- function(allScore){ return(allScore < 0.05)} # function that returns TRUE/FALSE for p-values<0.05
allGO2genes <- annFUN.org(whichOnto="BP", feasibleGenes=NULL, mapping="org.Hs.eg.db", ID="ensembl")Loading required package: org.Hs.eg.dbGOdata <- new("topGOdata",
ontology="BP",
allGenes=genes,
annot=annFUN.GO2genes,
GO2genes=allGO2genes,
geneSel=selection,
nodeSize=10)
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )results.ks <- runTest(GOdata, algorithm="classic", statistic="ks")
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasinggoEnrichment <- GenTable(GOdata, KS=results.ks, orderBy="KS", topNodes=20)
goEnrichment <- goEnrichment %>%
mutate(KS = ifelse(grepl("<", KS), 1e-30, KS))
goEnrichment$KS <- as.numeric(goEnrichment$KS)
goEnrichment <- goEnrichment[goEnrichment$KS<0.05,]
goEnrichment <- goEnrichment[,c("GO.ID","Term","KS")]
goEnrichment$Term <- gsub(" [a-z]*\\.\\.\\.$", "", goEnrichment$Term)
goEnrichment$Term <- gsub("\\.\\.\\.$", "", goEnrichment$Term)
goEnrichment$Term <- paste(goEnrichment$GO.ID, goEnrichment$Term, sep=", ")
goEnrichment$Term <- factor(goEnrichment$Term, levels=rev(goEnrichment$Term))
goEnrichment$KS <- as.numeric(goEnrichment$KS)
ggplot(goEnrichment, aes(x=Term, y=-log10(KS))) +
stat_summary(geom = "bar", fun = mean, position = "dodge") +
xlab("Biological process") +
ylab("Enrichment") +
ggtitle("Title") +
scale_y_continuous(breaks = round(seq(0, max(-log10(goEnrichment$KS)), by = 2), 1)) +
theme_bw(base_size=24) +
theme(
legend.position='none',
legend.background=element_rect(),
plot.title=element_text(angle=0, size=24, face="bold", vjust=1),
axis.text.x=element_text(angle=0, size=18, face="bold", hjust=1.10),
axis.text.y=element_text(angle=0, size=18, face="bold", vjust=0.5),
axis.title=element_text(size=24, face="bold"),
legend.key=element_blank(), #removes the border
legend.key.size=unit(1, "cm"), #Sets overall area/size of the legend
legend.text=element_text(size=18), #Text size
title=element_text(size=18)) +
guides(colour=guide_legend(override.aes=list(size=2.5))) +
coord_flip()
| Version | Author | Date |
|---|---|---|
| 8542407 | Anthony Hung | 2021-01-21 |
Additional analysis permuting samples:
Randomly shuffle the strain/unstrain labels 10 times and perform DE analysis on each, looking at the enriched GO terms to make sure the DE genes are not primarily driven by increased expression of certain genes in my cell type(s). Store the DE results to look at enrichments in OA gene sets later as well.
#scramble treatment labels (x3) and store the DE genes (each time, randomly assign 9 samples to be strain)
permuted_results <- vector("list", 12)
for(time in 1:10){
#set up DE and scramble labels
anno <- pData(RUVsOut)
x <- paste0(anno$Individual, anno$treatment)
anno$LibraryPrepBatch <- factor(anno$LibraryPrepBatch, levels = c("1", "2"))
anno$Replicate <- factor(anno$Replicate, levels = c("1", "2", "3"))
strain_indices <- sample(nrow(anno), 9)
anno$treatment <- "Unstrain"
anno$treatment[strain_indices] <- "Strain"
design <- model.matrix(~treatment + Individual + W_1 + W_2 + RIN,
data = anno)
colnames(design) <- gsub("treatment", "", colnames(design))
permuted_results[[time]] <- anno
#perform DE
#TMM Normalization
y <- DGEList(filt_counts)
y <- calcNormFactors(y, method = "TMM")
#Voom for differential expression
v1 <- voom(y, design)
corfit1 <- duplicateCorrelation(v1, design, block = x)
corfit1$consensus.correlation
v2 <- voom(y, design, block = x, correlation = corfit1$consensus.correlation)
corfit2 <- duplicateCorrelation(v2, design, block = x)
corfit2$consensus.correlation
fit <- lmFit(v2, design, block = x,
correlation = corfit2$consensus.correlation)
fit <- eBayes(fit)
tests <- colnames(fit$coefficients)
results <- vector(length = length(tests), mode = "list")
names(results) <- tests
for (test in tests[c(1:7)]) {
# Extract limma results
results[[test]] <- get_results(fit, coef = test)
# Add mutliple testing correction with ASH
output_ash <- run_ash(fit, coef = test)$result
results[[test]] <- cbind(results[[test]], lfsr = output_ash$lfsr,
lfdr = output_ash$lfdr, qvalue = output_ash$qvalue,
svalue = output_ash$svalue)
}
#store results
permuted_results[[10+time]] <- results[["Unstrain"]]
}Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable
Warning in glmgam.fit(dx, dy, coef.start = start, tol = tol, maxit =
maxit, : Too much damping - convergence tolerance not achievable#visualize what the permutations look like (what the scrambled labels look like)
for(time in 1:10){
print(permuted_results[[time]])
} Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Strain 10.0
18856_3_U 18856_3_U NA18856 M 3 Strain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Unstrain 10.0
18856_2_S 18856_2_S NA18856 M 2 Strain 9.6
19160_3_U 19160_3_U NA19160 M 3 Strain 10.0
18855_2_U 18855_2_U NA18855 F 2 Unstrain 10.0
19160_2_S 19160_2_S NA19160 M 2 Unstrain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Unstrain 8.5
19160_1_S 19160_1_S NA19160 M 1 Strain 9.9
19160_2_U 19160_2_U NA19160 M 2 Strain 10.0
18855_1_U 18855_1_U NA18855 F 1 Strain 10.0
18856_3_S 18856_3_S NA18856 M 3 Strain 9.6
18856_2_U 18856_2_U NA18856 M 2 Strain 9.5
18855_3_U 18855_3_U NA18855 F 3 Unstrain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Strain 10.0
19160_3_S 19160_3_S NA19160 M 3 Unstrain 10.0
18856_3_U 18856_3_U NA18856 M 3 Strain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Strain 10.0
18856_2_S 18856_2_S NA18856 M 2 Strain 9.6
19160_3_U 19160_3_U NA19160 M 3 Unstrain 10.0
18855_2_U 18855_2_U NA18855 F 2 Strain 10.0
19160_2_S 19160_2_S NA19160 M 2 Strain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Strain 8.5
19160_1_S 19160_1_S NA19160 M 1 Unstrain 9.9
19160_2_U 19160_2_U NA19160 M 2 Unstrain 10.0
18855_1_U 18855_1_U NA18855 F 1 Unstrain 10.0
18856_3_S 18856_3_S NA18856 M 3 Unstrain 9.6
18856_2_U 18856_2_U NA18856 M 2 Strain 9.5
18855_3_U 18855_3_U NA18855 F 3 Strain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Unstrain 10.0
18856_3_U 18856_3_U NA18856 M 3 Strain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Strain 10.0
18856_2_S 18856_2_S NA18856 M 2 Unstrain 9.6
19160_3_U 19160_3_U NA19160 M 3 Strain 10.0
18855_2_U 18855_2_U NA18855 F 2 Strain 10.0
19160_2_S 19160_2_S NA19160 M 2 Strain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Unstrain 8.5
19160_1_S 19160_1_S NA19160 M 1 Strain 9.9
19160_2_U 19160_2_U NA19160 M 2 Strain 10.0
18855_1_U 18855_1_U NA18855 F 1 Unstrain 10.0
18856_3_S 18856_3_S NA18856 M 3 Strain 9.6
18856_2_U 18856_2_U NA18856 M 2 Unstrain 9.5
18855_3_U 18855_3_U NA18855 F 3 Strain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Strain 10.0
18856_3_U 18856_3_U NA18856 M 3 Unstrain 10.0
18856_1_U 18856_1_U NA18856 M 1 Strain 9.9
18855_2_S 18855_2_S NA18855 F 2 Unstrain 10.0
18856_2_S 18856_2_S NA18856 M 2 Strain 9.6
19160_3_U 19160_3_U NA19160 M 3 Strain 10.0
18855_2_U 18855_2_U NA18855 F 2 Unstrain 10.0
19160_2_S 19160_2_S NA19160 M 2 Strain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Strain 8.5
19160_1_S 19160_1_S NA19160 M 1 Strain 9.9
19160_2_U 19160_2_U NA19160 M 2 Strain 10.0
18855_1_U 18855_1_U NA18855 F 1 Unstrain 10.0
18856_3_S 18856_3_S NA18856 M 3 Unstrain 9.6
18856_2_U 18856_2_U NA18856 M 2 Strain 9.5
18855_3_U 18855_3_U NA18855 F 3 Unstrain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Unstrain 10.0
18856_3_U 18856_3_U NA18856 M 3 Strain 10.0
18856_1_U 18856_1_U NA18856 M 1 Strain 9.9
18855_2_S 18855_2_S NA18855 F 2 Unstrain 10.0
18856_2_S 18856_2_S NA18856 M 2 Strain 9.6
19160_3_U 19160_3_U NA19160 M 3 Unstrain 10.0
18855_2_U 18855_2_U NA18855 F 2 Strain 10.0
19160_2_S 19160_2_S NA19160 M 2 Strain 10.0
18855_1_S 18855_1_S NA18855 F 1 Strain 9.9
18856_1_S 18856_1_S NA18856 M 1 Strain 8.5
19160_1_S 19160_1_S NA19160 M 1 Strain 9.9
19160_2_U 19160_2_U NA19160 M 2 Unstrain 10.0
18855_1_U 18855_1_U NA18855 F 1 Unstrain 10.0
18856_3_S 18856_3_S NA18856 M 3 Unstrain 9.6
18856_2_U 18856_2_U NA18856 M 2 Unstrain 9.5
18855_3_U 18855_3_U NA18855 F 3 Strain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Strain 10.0
18856_3_U 18856_3_U NA18856 M 3 Strain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Unstrain 10.0
18856_2_S 18856_2_S NA18856 M 2 Unstrain 9.6
19160_3_U 19160_3_U NA19160 M 3 Unstrain 10.0
18855_2_U 18855_2_U NA18855 F 2 Unstrain 10.0
19160_2_S 19160_2_S NA19160 M 2 Strain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Strain 8.5
19160_1_S 19160_1_S NA19160 M 1 Strain 9.9
19160_2_U 19160_2_U NA19160 M 2 Strain 10.0
18855_1_U 18855_1_U NA18855 F 1 Strain 10.0
18856_3_S 18856_3_S NA18856 M 3 Strain 9.6
18856_2_U 18856_2_U NA18856 M 2 Unstrain 9.5
18855_3_U 18855_3_U NA18855 F 3 Strain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Unstrain 10.0
18856_3_U 18856_3_U NA18856 M 3 Strain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Strain 10.0
18856_2_S 18856_2_S NA18856 M 2 Strain 9.6
19160_3_U 19160_3_U NA19160 M 3 Strain 10.0
18855_2_U 18855_2_U NA18855 F 2 Unstrain 10.0
19160_2_S 19160_2_S NA19160 M 2 Unstrain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Strain 8.5
19160_1_S 19160_1_S NA19160 M 1 Strain 9.9
19160_2_U 19160_2_U NA19160 M 2 Unstrain 10.0
18855_1_U 18855_1_U NA18855 F 1 Strain 10.0
18856_3_S 18856_3_S NA18856 M 3 Unstrain 9.6
18856_2_U 18856_2_U NA18856 M 2 Strain 9.5
18855_3_U 18855_3_U NA18855 F 3 Strain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Unstrain 10.0
18856_3_U 18856_3_U NA18856 M 3 Strain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Strain 10.0
18856_2_S 18856_2_S NA18856 M 2 Unstrain 9.6
19160_3_U 19160_3_U NA19160 M 3 Unstrain 10.0
18855_2_U 18855_2_U NA18855 F 2 Strain 10.0
19160_2_S 19160_2_S NA19160 M 2 Unstrain 10.0
18855_1_S 18855_1_S NA18855 F 1 Strain 9.9
18856_1_S 18856_1_S NA18856 M 1 Strain 8.5
19160_1_S 19160_1_S NA19160 M 1 Strain 9.9
19160_2_U 19160_2_U NA19160 M 2 Strain 10.0
18855_1_U 18855_1_U NA18855 F 1 Strain 10.0
18856_3_S 18856_3_S NA18856 M 3 Unstrain 9.6
18856_2_U 18856_2_U NA18856 M 2 Unstrain 9.5
18855_3_U 18855_3_U NA18855 F 3 Strain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Unstrain 10.0
19160_3_S 19160_3_S NA19160 M 3 Unstrain 10.0
18856_3_U 18856_3_U NA18856 M 3 Unstrain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Strain 10.0
18856_2_S 18856_2_S NA18856 M 2 Strain 9.6
19160_3_U 19160_3_U NA19160 M 3 Strain 10.0
18855_2_U 18855_2_U NA18855 F 2 Unstrain 10.0
19160_2_S 19160_2_S NA19160 M 2 Strain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Strain 8.5
19160_1_S 19160_1_S NA19160 M 1 Unstrain 9.9
19160_2_U 19160_2_U NA19160 M 2 Strain 10.0
18855_1_U 18855_1_U NA18855 F 1 Strain 10.0
18856_3_S 18856_3_S NA18856 M 3 Strain 9.6
18856_2_U 18856_2_U NA18856 M 2 Strain 9.5
18855_3_U 18855_3_U NA18855 F 3 Unstrain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497
Sample_ID Individual Sex Replicate treatment RIN
18855_3_S 18855_3_S NA18855 F 3 Strain 10.0
19160_3_S 19160_3_S NA19160 M 3 Strain 10.0
18856_3_U 18856_3_U NA18856 M 3 Unstrain 10.0
18856_1_U 18856_1_U NA18856 M 1 Unstrain 9.9
18855_2_S 18855_2_S NA18855 F 2 Strain 10.0
18856_2_S 18856_2_S NA18856 M 2 Unstrain 9.6
19160_3_U 19160_3_U NA19160 M 3 Strain 10.0
18855_2_U 18855_2_U NA18855 F 2 Strain 10.0
19160_2_S 19160_2_S NA19160 M 2 Unstrain 10.0
18855_1_S 18855_1_S NA18855 F 1 Unstrain 9.9
18856_1_S 18856_1_S NA18856 M 1 Unstrain 8.5
19160_1_S 19160_1_S NA19160 M 1 Unstrain 9.9
19160_2_U 19160_2_U NA19160 M 2 Unstrain 10.0
18855_1_U 18855_1_U NA18855 F 1 Strain 10.0
18856_3_S 18856_3_S NA18856 M 3 Strain 9.6
18856_2_U 18856_2_U NA18856 M 2 Strain 9.5
18855_3_U 18855_3_U NA18855 F 3 Strain 10.0
LibraryPrepBatch LibSize W_1 W_2
18855_3_S 2 15419569 0.541963415 -1.1181740
19160_3_S 2 15177653 0.376349314 -1.4100700
18856_3_U 2 13722909 0.857248982 -0.8808562
18856_1_U 1 18647599 1.113674153 -1.5466409
18855_2_S 1 18256163 0.733375747 -0.7617020
18856_2_S 1 16782764 -0.051863231 -1.3484210
19160_3_U 2 19839604 0.224499121 -1.5560509
18855_2_U 1 18618691 0.665015743 -0.9464050
19160_2_S 1 16622407 0.308745015 -1.5149287
18855_1_S 1 19712744 0.626769726 -1.5366695
18856_1_S 1 13571439 1.038716806 -1.2664724
19160_1_S 1 20979958 0.461484490 -2.0300296
19160_2_U 1 20924281 0.002654243 -1.9184820
18855_1_U 1 20146697 0.649711618 -1.4110342
18856_3_S 2 15807863 0.728747075 -0.9909086
18856_2_U 1 15142152 0.359175096 -1.0809905
18855_3_U 2 18157918 0.487681866 -1.2290497Perform ks test for enrichments of GO terms amongst the permuted DE genes
for (permutation in 11:20){
results <- permuted_results[[permutation]]
# k-s test making use of ranked based on p-values
genes <- results$qvalue
names(genes) <- rownames(results)
selection <- function(allScore){ return(allScore < 0.05)} # function that returns TRUE/FALSE for p-values<0.05
allGO2genes <- annFUN.org(whichOnto="BP", feasibleGenes=NULL, mapping="org.Hs.eg.db", ID="ensembl")
GOdata <- new("topGOdata",
ontology="BP",
allGenes=genes,
annot=annFUN.GO2genes,
GO2genes=allGO2genes,
geneSel=selection,
nodeSize=10)
results.ks <- runTest(GOdata, algorithm="classic", statistic="ks")
goEnrichment <- GenTable(GOdata, KS=results.ks, orderBy="KS", topNodes=20)
goEnrichment <- goEnrichment %>%
mutate(KS = ifelse(grepl("<", KS), 1e-30, KS))
goEnrichment$KS <- as.numeric(goEnrichment$KS)
goEnrichment <- goEnrichment[goEnrichment$KS<0.05,]
goEnrichment <- goEnrichment[,c("GO.ID","Term","KS")]
goEnrichment$Term <- gsub(" [a-z]*\\.\\.\\.$", "", goEnrichment$Term)
goEnrichment$Term <- gsub("\\.\\.\\.$", "", goEnrichment$Term)
goEnrichment$Term <- paste(goEnrichment$GO.ID, goEnrichment$Term, sep=", ")
goEnrichment$Term <- factor(goEnrichment$Term, levels=rev(goEnrichment$Term))
goEnrichment$KS <- as.numeric(goEnrichment$KS)
print(goEnrichment$Term)
}
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0010591, regulation of lamellipodium assembly
[2] GO:0043038, amino acid activation
[3] GO:0043039, tRNA aminoacylation
[4] GO:0003230, cardiac atrium development
[5] GO:0006418, tRNA aminoacylation for protein
[6] GO:0010592, positive regulation of lamellipodium
[7] GO:0032635, interleukin-6 production
[8] GO:0043489, RNA stabilization
[9] GO:0060021, roof of mouth development
[10] GO:0021536, diencephalon development
[11] GO:0003209, cardiac atrium morphogenesis
[12] GO:0003283, atrial septum development
[13] GO:0002820, negative regulation of adaptive immune
[14] GO:0050919, negative chemotaxis
[15] GO:0006928, movement of cell or subcellular
[16] GO:0048255, mRNA stabilization
[17] GO:0045669, positive regulation of osteoblast
[18] GO:0061029, eyelid development in camera-type eye
[19] GO:0043486, histone exchange
[20] GO:0010950, positive regulation of endopeptidase
20 Levels: GO:0010950, positive regulation of endopeptidase ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0007186, G protein-coupled receptor signaling
[2] GO:0032501, multicellular organismal process
[3] GO:0003008, system process
[4] GO:0048856, anatomical structure development
[5] GO:0061138, morphogenesis of a branching epithelium
[6] GO:0009888, tissue development
[7] GO:0001763, morphogenesis of a branching structure
[8] GO:0048513, animal organ development
[9] GO:0032502, developmental process
[10] GO:0009653, anatomical structure morphogenesis
[11] GO:0001934, positive regulation of protein
[12] GO:0007275, multicellular organism development
[13] GO:0051239, regulation of multicellular organismal
[14] GO:0048731, system development
[15] GO:0022612, gland morphogenesis
[16] GO:0042327, positive regulation of phosphorylation
[17] GO:0048645, animal organ formation
[18] GO:0051241, negative regulation of multicellular
[19] GO:0030154, cell differentiation
[20] GO:0048646, anatomical structure formation involved
20 Levels: GO:0048646, anatomical structure formation involved ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0010591, regulation of lamellipodium assembly
[2] GO:0043038, amino acid activation
[3] GO:0043039, tRNA aminoacylation
[4] GO:0003230, cardiac atrium development
[5] GO:0006418, tRNA aminoacylation for protein
[6] GO:0010592, positive regulation of lamellipodium
[7] GO:0032635, interleukin-6 production
[8] GO:0043489, RNA stabilization
[9] GO:0060021, roof of mouth development
[10] GO:0021536, diencephalon development
[11] GO:0003209, cardiac atrium morphogenesis
[12] GO:0003283, atrial septum development
[13] GO:0002820, negative regulation of adaptive immune
[14] GO:0050919, negative chemotaxis
[15] GO:0006928, movement of cell or subcellular
[16] GO:0048255, mRNA stabilization
[17] GO:0045669, positive regulation of osteoblast
[18] GO:0061029, eyelid development in camera-type eye
[19] GO:0043486, histone exchange
[20] GO:0010950, positive regulation of endopeptidase
20 Levels: GO:0010950, positive regulation of endopeptidase ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0006928, movement of cell or subcellular
[2] GO:0007166, cell surface receptor signaling pathway
[3] GO:0040011, locomotion
[4] GO:0042127, regulation of cell proliferation
[5] GO:0007165, signal transduction
[6] GO:0048731, system development
[7] GO:0023052, signaling
[8] GO:0009653, anatomical structure morphogenesis
[9] GO:0030154, cell differentiation
[10] GO:0007154, cell communication
[11] GO:0048869, cellular developmental process
[12] GO:0050896, response to stimulus
[13] GO:0009605, response to external stimulus
[14] GO:0051270, regulation of cellular component
[15] GO:0030334, regulation of cell migration
[16] GO:0016477, cell migration
[17] GO:0051239, regulation of multicellular organismal
[18] GO:0048870, cell motility
[19] GO:0051674, localization of cell
[20] GO:0051716, cellular response to stimulus
20 Levels: GO:0051716, cellular response to stimulus ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0010591, regulation of lamellipodium assembly
[2] GO:0043038, amino acid activation
[3] GO:0043039, tRNA aminoacylation
[4] GO:0003230, cardiac atrium development
[5] GO:0006418, tRNA aminoacylation for protein
[6] GO:0010592, positive regulation of lamellipodium
[7] GO:0032635, interleukin-6 production
[8] GO:0043489, RNA stabilization
[9] GO:0060021, roof of mouth development
[10] GO:0021536, diencephalon development
[11] GO:0003209, cardiac atrium morphogenesis
[12] GO:0003283, atrial septum development
[13] GO:0002820, negative regulation of adaptive immune
[14] GO:0050919, negative chemotaxis
[15] GO:0006928, movement of cell or subcellular
[16] GO:0048255, mRNA stabilization
[17] GO:0045669, positive regulation of osteoblast
[18] GO:0061029, eyelid development in camera-type eye
[19] GO:0043486, histone exchange
[20] GO:0010950, positive regulation of endopeptidase
20 Levels: GO:0010950, positive regulation of endopeptidase ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0010591, regulation of lamellipodium assembly
[2] GO:0043038, amino acid activation
[3] GO:0043039, tRNA aminoacylation
[4] GO:0003230, cardiac atrium development
[5] GO:0006418, tRNA aminoacylation for protein
[6] GO:0010592, positive regulation of lamellipodium
[7] GO:0032635, interleukin-6 production
[8] GO:0043489, RNA stabilization
[9] GO:0060021, roof of mouth development
[10] GO:0021536, diencephalon development
[11] GO:0003209, cardiac atrium morphogenesis
[12] GO:0003283, atrial septum development
[13] GO:0002820, negative regulation of adaptive immune
[14] GO:0050919, negative chemotaxis
[15] GO:0006928, movement of cell or subcellular
[16] GO:0048255, mRNA stabilization
[17] GO:0045669, positive regulation of osteoblast
[18] GO:0061029, eyelid development in camera-type eye
[19] GO:0043486, histone exchange
[20] GO:0010950, positive regulation of endopeptidase
20 Levels: GO:0010950, positive regulation of endopeptidase ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0006614, SRP-dependent cotranslational protein
[2] GO:0045047, protein targeting to ER
[3] GO:0072599, establishment of protein localization
[4] GO:0006613, cotranslational protein targeting to
[5] GO:0000184, nuclear-transcribed mRNA catabolic
[6] GO:0070972, protein localization to endoplasmic
[7] GO:0019083, viral transcription
[8] GO:0019080, viral gene expression
[9] GO:0006612, protein targeting to membrane
[10] GO:0006413, translational initiation
[11] GO:0000956, nuclear-transcribed mRNA catabolic
[12] GO:0006605, protein targeting
[13] GO:0006412, translation
[14] GO:0043043, peptide biosynthetic process
[15] GO:0010469, regulation of signaling receptor
[16] GO:0030198, extracellular matrix organization
[17] GO:0006518, peptide metabolic process
[18] GO:0035295, tube development
[19] GO:0043062, extracellular structure organization
[20] GO:1901342, regulation of vasculature development
20 Levels: GO:1901342, regulation of vasculature development ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0010591, regulation of lamellipodium assembly
[2] GO:0043038, amino acid activation
[3] GO:0043039, tRNA aminoacylation
[4] GO:0003230, cardiac atrium development
[5] GO:0006418, tRNA aminoacylation for protein
[6] GO:0010592, positive regulation of lamellipodium
[7] GO:0032635, interleukin-6 production
[8] GO:0043489, RNA stabilization
[9] GO:0060021, roof of mouth development
[10] GO:0021536, diencephalon development
[11] GO:0003209, cardiac atrium morphogenesis
[12] GO:0003283, atrial septum development
[13] GO:0002820, negative regulation of adaptive immune
[14] GO:0050919, negative chemotaxis
[15] GO:0006928, movement of cell or subcellular
[16] GO:0048255, mRNA stabilization
[17] GO:0045669, positive regulation of osteoblast
[18] GO:0061029, eyelid development in camera-type eye
[19] GO:0043486, histone exchange
[20] GO:0010950, positive regulation of endopeptidase
20 Levels: GO:0010950, positive regulation of endopeptidase ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0010591, regulation of lamellipodium assembly
[2] GO:0043038, amino acid activation
[3] GO:0043039, tRNA aminoacylation
[4] GO:0003230, cardiac atrium development
[5] GO:0006418, tRNA aminoacylation for protein
[6] GO:0010592, positive regulation of lamellipodium
[7] GO:0032635, interleukin-6 production
[8] GO:0043489, RNA stabilization
[9] GO:0060021, roof of mouth development
[10] GO:0021536, diencephalon development
[11] GO:0003209, cardiac atrium morphogenesis
[12] GO:0003283, atrial septum development
[13] GO:0002820, negative regulation of adaptive immune
[14] GO:0050919, negative chemotaxis
[15] GO:0006928, movement of cell or subcellular
[16] GO:0048255, mRNA stabilization
[17] GO:0045669, positive regulation of osteoblast
[18] GO:0061029, eyelid development in camera-type eye
[19] GO:0043486, histone exchange
[20] GO:0010950, positive regulation of endopeptidase
20 Levels: GO:0010950, positive regulation of endopeptidase ...
Building most specific GOs ..... ( 9780 GO terms found. )
Build GO DAG topology .......... ( 13861 GO terms and 32652 relations. )
Annotating nodes ............... ( 9204 genes annotated to the GO terms. )
-- Classic Algorithm --
the algorithm is scoring 5178 nontrivial nodes
parameters:
test statistic: ks
score order: increasing [1] GO:0010591, regulation of lamellipodium assembly
[2] GO:0043038, amino acid activation
[3] GO:0043039, tRNA aminoacylation
[4] GO:0003230, cardiac atrium development
[5] GO:0006418, tRNA aminoacylation for protein
[6] GO:0010592, positive regulation of lamellipodium
[7] GO:0032635, interleukin-6 production
[8] GO:0043489, RNA stabilization
[9] GO:0060021, roof of mouth development
[10] GO:0021536, diencephalon development
[11] GO:0003209, cardiac atrium morphogenesis
[12] GO:0003283, atrial septum development
[13] GO:0002820, negative regulation of adaptive immune
[14] GO:0050919, negative chemotaxis
[15] GO:0006928, movement of cell or subcellular
[16] GO:0048255, mRNA stabilization
[17] GO:0045669, positive regulation of osteoblast
[18] GO:0061029, eyelid development in camera-type eye
[19] GO:0043486, histone exchange
[20] GO:0010950, positive regulation of endopeptidase
20 Levels: GO:0010950, positive regulation of endopeptidase ...Enrichment Function
Use fisher’s exact test to compute enrichment of DE genes amongst the OA related gene sets.
enrich <- function(genes_of_interest, DEgenes, backgroundgenes){
#fill contingency table
DE_interest <- length(intersect(genes_of_interest, DEgenes))
background_interest <- length(intersect(genes_of_interest, backgroundgenes))
DE_non_interest <- length(DEgenes) - DE_interest
background_non_interest <- length(backgroundgenes) - background_interest
contingency_table <- matrix(c(DE_interest, background_interest, DE_non_interest, background_non_interest), nrow = 2, ncol = 2, byrow = TRUE)
#perform test
print(contingency_table)
return(fisher.test(contingency_table, alternative="greater"))
}
enrich(fungenomic, DE_genes_ensg, other_genes_ensg) [,1] [,2]
[1,] 49 293
[2,] 938 9206
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 0.001826
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
1.242775 Inf
sample estimates:
odds ratio
1.641232 enrich(cross_omic, DE_genes_ensg, other_genes_ensg) [,1] [,2]
[1,] 42 296
[2,] 945 9203
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 0.03712
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
1.025732 Inf
sample estimates:
odds ratio
1.381779 Look at enrichments from permuted DE analysis (scrambled strain labels to make sure the FET results aren’t just because some genes are more likely to be detected as DE in these cells)
threshold <- 0.05
for(permutation in 11:20){
print(permutation)
DE_results <- permuted_results[[permutation]]
DE_genes_ensg <- rownames(DE_results)[DE_results$qvalue < threshold]
DE_genes <- gene_anno$external_gene_name[match(DE_genes_ensg, gene_anno$ensembl_gene_id)]
other_genes_ensg <- rownames(DE_results)[DE_results$qvalue >= threshold]
other_genes <- gene_anno$external_gene_name[match(other_genes_ensg, gene_anno$ensembl_gene_id)]
print(enrich(fungenomic, DE_genes_ensg, other_genes_ensg))
print(enrich(cross_omic, DE_genes_ensg, other_genes_ensg))
}[1] 11
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 12
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 13
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 14
[,1] [,2]
[1,] 0 342
[2,] 7 10137
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 7 10141
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 15
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 16
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 17
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 18
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 19
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[1] 20
[,1] [,2]
[1,] 0 342
[2,] 0 10144
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
[,1] [,2]
[1,] 0 338
[2,] 0 10148
Fisher's Exact Test for Count Data
data: contingency_table
p-value = 1
alternative hypothesis: true odds ratio is greater than 1
95 percent confidence interval:
0 Inf
sample estimates:
odds ratio
0
sessionInfo()R version 3.6.1 (2019-07-05)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Scientific Linux 7.4 (Nitrogen)
Matrix products: default
BLAS/LAPACK: /software/openblas-0.2.19-el7-x86_64/lib/libopenblas_haswellp-r0.2.19.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats4 parallel stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] org.Hs.eg.db_3.8.2 topGO_2.36.0
[3] SparseM_1.78 GO.db_3.8.2
[5] AnnotationDbi_1.46.0 graph_1.62.0
[7] RUVSeq_1.18.0 EDASeq_2.18.0
[9] ShortRead_1.42.0 GenomicAlignments_1.20.1
[11] SummarizedExperiment_1.14.1 DelayedArray_0.10.0
[13] matrixStats_0.57.0 Rsamtools_2.0.0
[15] GenomicRanges_1.36.1 GenomeInfoDb_1.20.0
[17] Biostrings_2.52.0 XVector_0.24.0
[19] IRanges_2.18.3 S4Vectors_0.22.1
[21] BiocParallel_1.18.1 Biobase_2.44.0
[23] BiocGenerics_0.30.0 ggplot2_3.3.3
[25] ashr_2.2-47 tidyr_1.1.2
[27] edgeR_3.26.5 dplyr_1.0.2
[29] plyr_1.8.6 limma_3.40.6
loaded via a namespace (and not attached):
[1] bitops_1.0-6 fs_1.3.1 bit64_0.9-7
[4] httr_1.4.2 progress_1.2.2 RColorBrewer_1.1-2
[7] rprojroot_2.0.2 tools_3.6.1 R6_2.5.0
[10] irlba_2.3.3 DBI_1.1.0 colorspace_2.0-0
[13] withr_2.3.0 prettyunits_1.1.1 tidyselect_1.1.0
[16] bit_4.0.4 compiler_3.6.1 git2r_0.26.1
[19] labeling_0.4.2 rtracklayer_1.44.0 scales_1.1.1
[22] SQUAREM_2020.4 genefilter_1.66.0 DESeq_1.36.0
[25] mixsqp_0.3-43 stringr_1.4.0 digest_0.6.27
[28] rmarkdown_1.13 R.utils_2.9.0 pkgconfig_2.0.3
[31] htmltools_0.5.0 invgamma_1.1 rlang_0.4.10
[34] RSQLite_2.1.1 farver_2.0.3 generics_0.0.2
[37] hwriter_1.3.2 R.oo_1.24.0 RCurl_1.98-1.1
[40] magrittr_2.0.1 GenomeInfoDbData_1.2.1 Matrix_1.2-18
[43] Rcpp_1.0.5 munsell_0.5.0 etrunct_0.1
[46] lifecycle_0.2.0 R.methodsS3_1.8.1 stringi_1.4.6
[49] whisker_0.3-2 yaml_2.2.1 MASS_7.3-52
[52] zlibbioc_1.30.0 grid_3.6.1 blob_1.2.0
[55] promises_1.1.1 crayon_1.3.4 lattice_0.20-41
[58] splines_3.6.1 GenomicFeatures_1.36.3 annotate_1.62.0
[61] hms_0.5.3 locfit_1.5-9.1 knitr_1.23
[64] pillar_1.4.7 biomaRt_2.40.1 geneplotter_1.62.0
[67] XML_3.98-1.20 glue_1.4.2 evaluate_0.14
[70] latticeExtra_0.6-28 vctrs_0.3.6 httpuv_1.5.1
[73] gtable_0.3.0 purrr_0.3.4 xfun_0.8
[76] aroma.light_3.14.0 xtable_1.8-4 later_1.1.0.1
[79] survival_2.44-1.1 truncnorm_1.0-8 tibble_3.0.4
[82] memoise_1.1.0 workflowr_1.6.2 statmod_1.4.32
[85] ellipsis_0.3.1